This kit is designed to make a reliable risk assessment for Microcystin/Nodalarin in swimming waters. The Microcystin Cyanotox qPCR detection kit is based on the fast, sensitive, and proven primers/probe qPCR technique. Microcystin and nodalarin are similar toxins with similar genes and are regarded as equivalent. The microcystin Cyanotox qPCR detection kit is designed to recognize both types of toxins at once. In this kit we use the name microcystin for both toxins. The used qPCR primers and probe recognizes the mcyJ gene. McyJ is a gene from the biochemical pathway for the production of microcystins and is only singular present in this pathway. This makes it a good candidate to determine the copy number of microcystin biochemical pathways in water samples.
The toxic gene potential (TGP) was determined with this kit. The toxic gene potential is defined as the amount of toxin that can been produced by one copy of the microcystin biochemical pathway. This value can be used to make a reliable risk assessment for Microcystin.
Detection as a service
We offer Microcystin Cyanotox qPCR detection also as a service. See our eDNA service website (in Dutch) or contact us for more information about sampling, sending, and pricing.
Toxin information
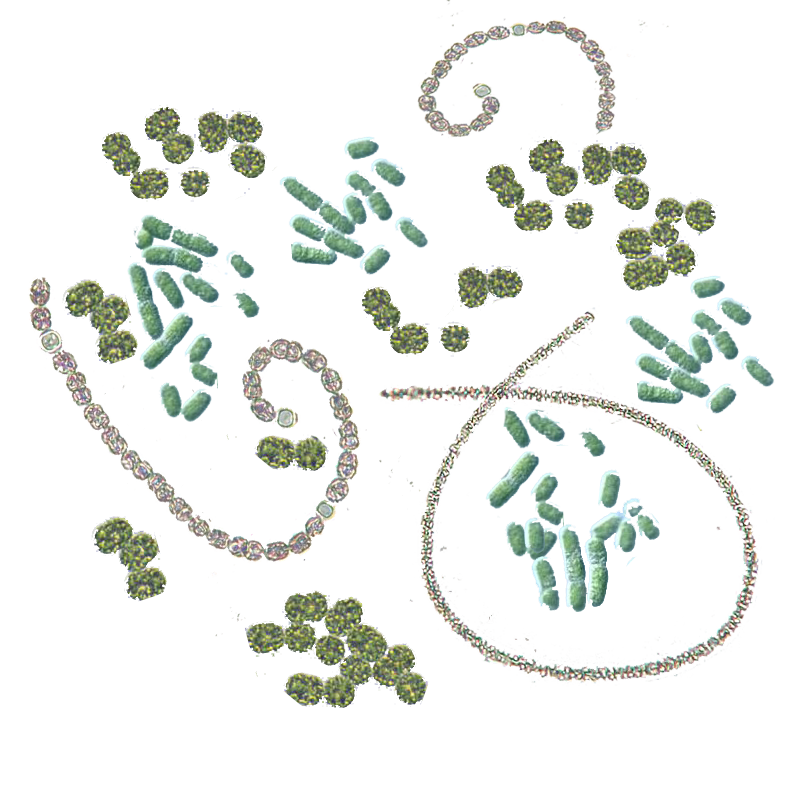
Microcystins — or cyanoginosins — are a class of toxins produced by certain freshwater cyanobacteria; primarily Microcystis aeruginosa but also other Microcystis, as well as members of the Planktothrix, Anabaena, Oscillatoria and Nostoc genera. Humans can be exposed to these toxins in numoures ways: swallowing, skin contact or inhaling contaminated water. Bloming of microcystin-producing bacteria can overwhelm the filter capacities of drinkingwater treatment plants. Some evidence shows the toxin can be transported by irrigation into the food chain. Microcystins are chemically stable over a wide range of temperature and pH, possibly as a result of their cyclic structure. (source: wikipedia)
Primer design
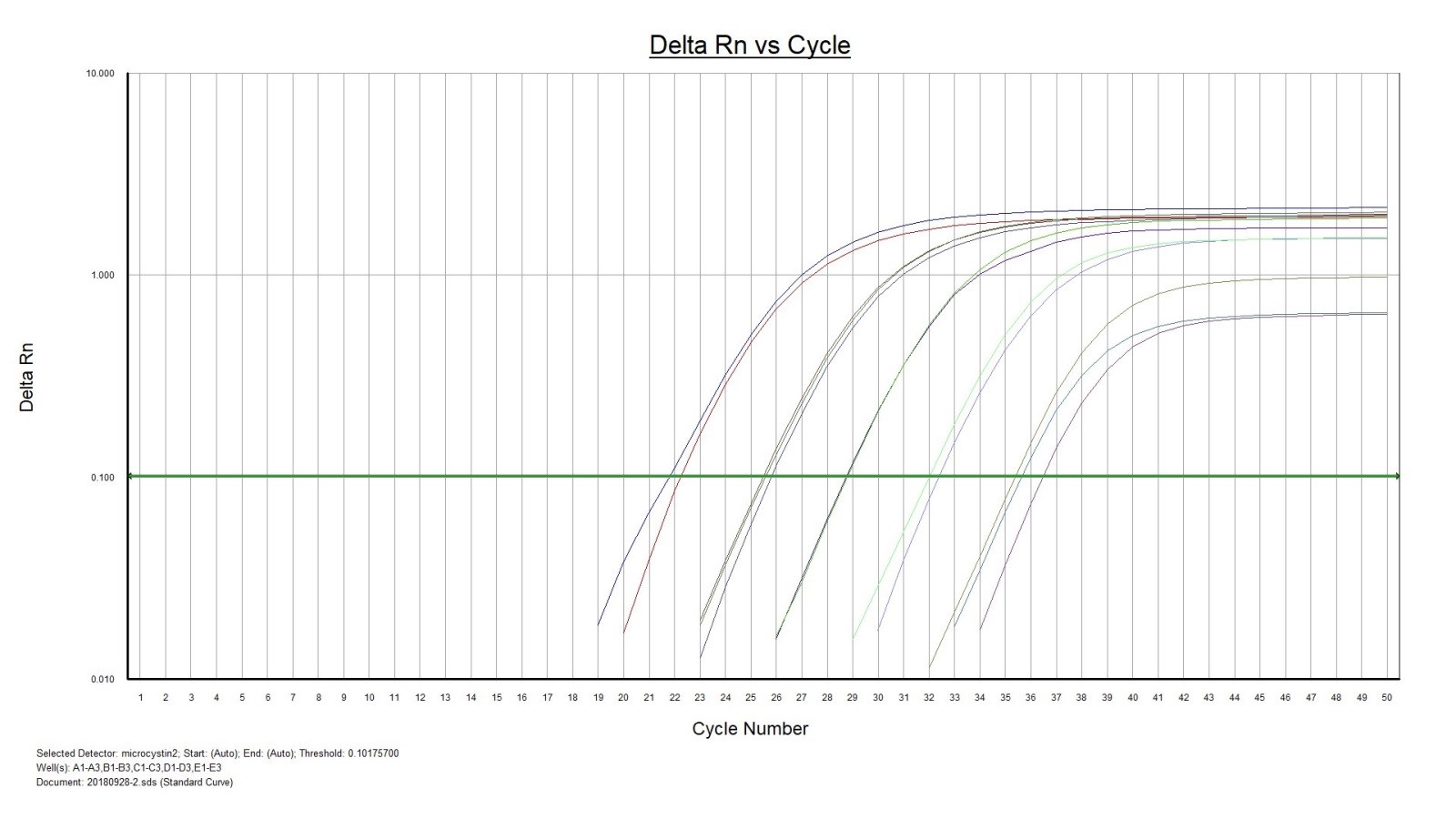
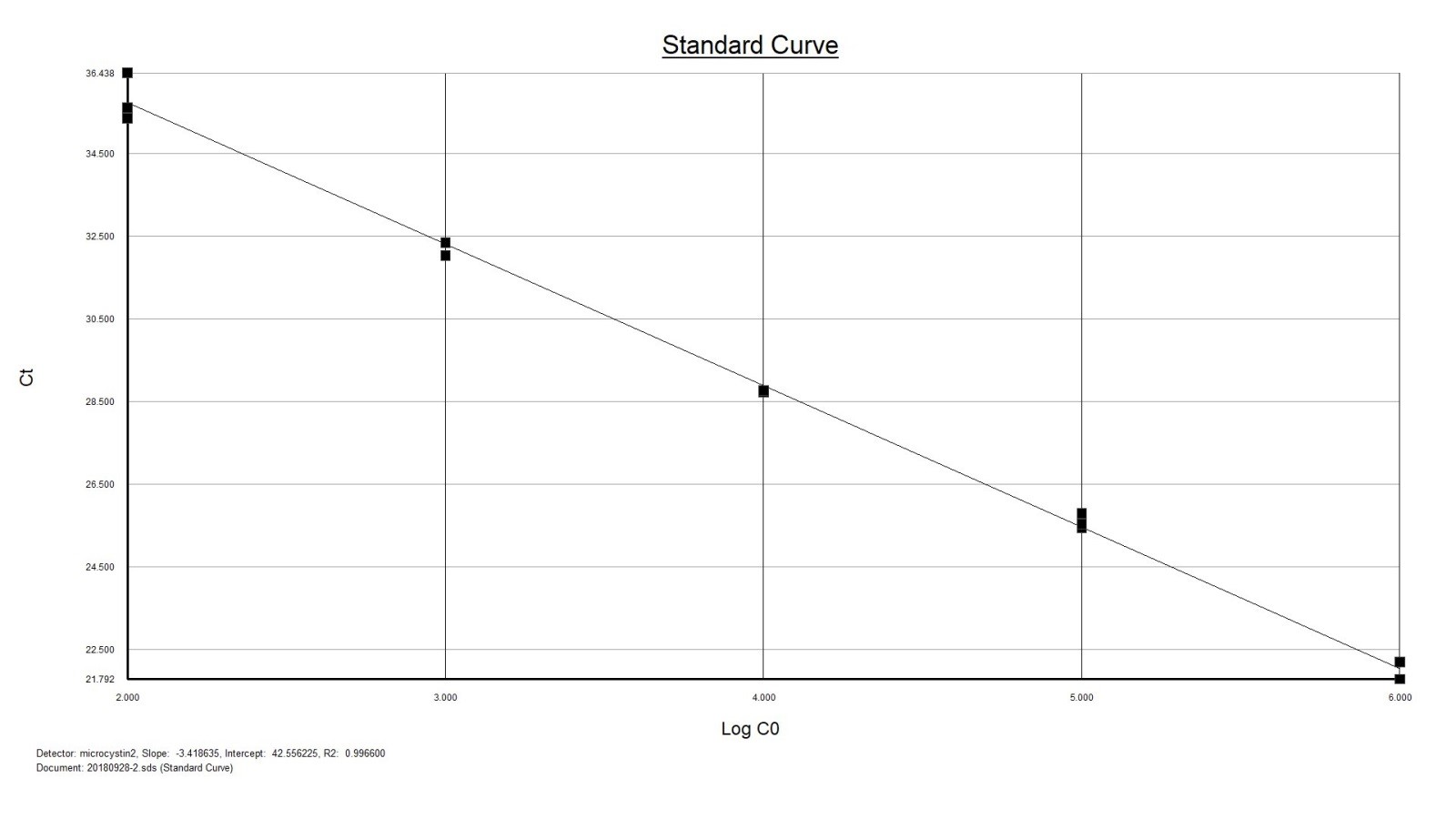
The primers and probe are specially designed to be used with eDNA samples and have the following properties:
- Highest possible sensitivity (<10 DNA copies per reaction).
- Strong fluorescence signal with low background noise. Isolated environmental samples contain residues of naturally occurring auto fluoresce substances that will interfere with the measurements. A strong fluorescence signal from the analyses is required for these kind of samples.
- 100% specificity. Isolated DNA from environmental samples contains billions of DNA fragments from bacteria, protozoa, plants, animals, etc.
The kit is developed and optimized to be used on eDNA isolates purified using the eDNA isolation kit (#SYL002) from Sylphium molecular ecology.
Other eDNA isolation methods/kits can be used as well. Please contact us to get more information how to use the obtained isolates from other methods/kits to get reliable results with the kit.
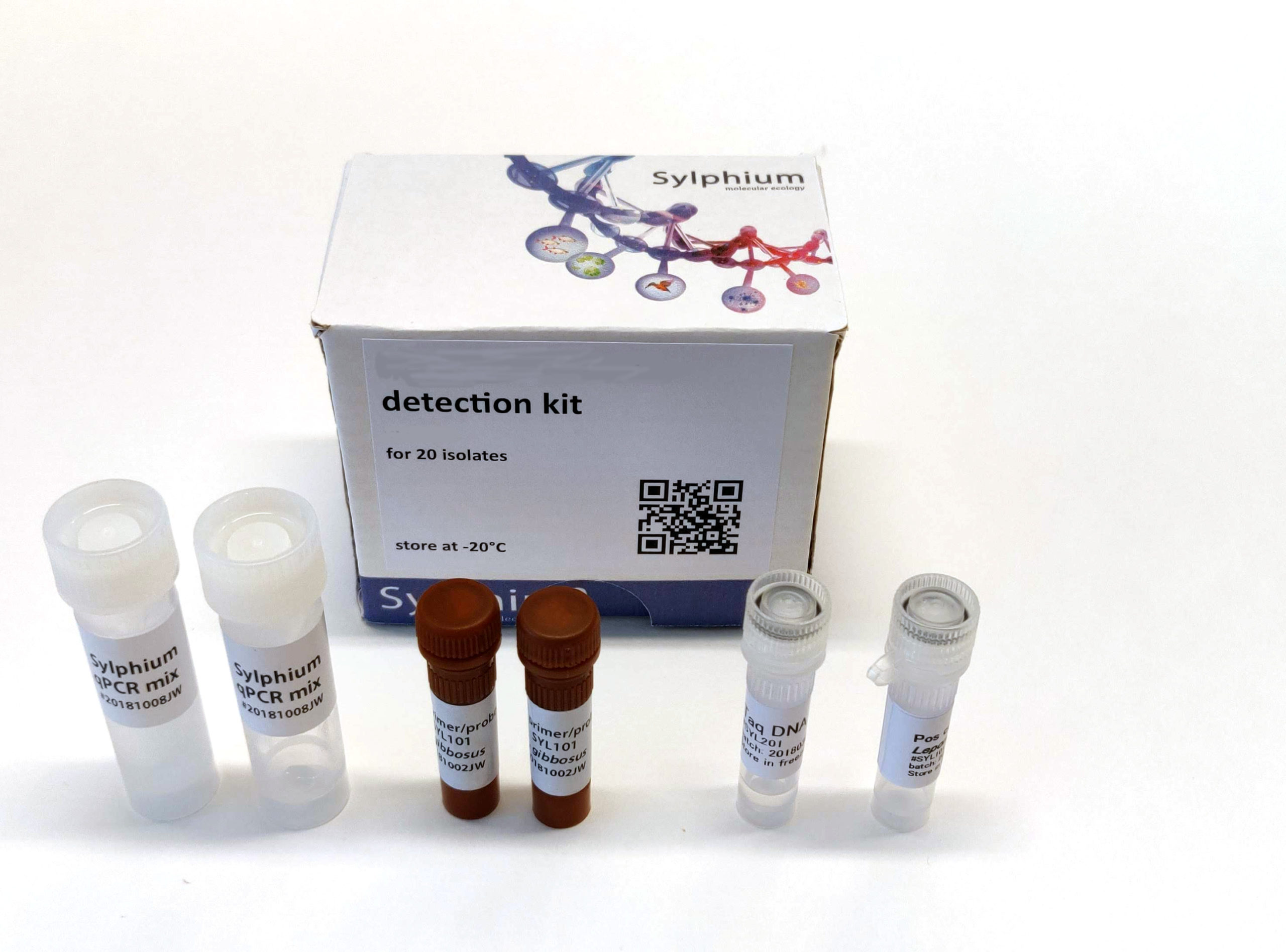
Kit contents
The kit contains materials for an 3-fold analyses on 54 samples including all necessary controls.
- Positive control (McyJ gene)
- 2x Sylphium qPCR mix (100 reactions) without primers and probes
- 2x Primer/probe mix (100 reactions) for detection of McyJ gene (FAM dye)
- 1x Taq DNA polymerase (200 reactions)
- Protocol and primer/probe validation report