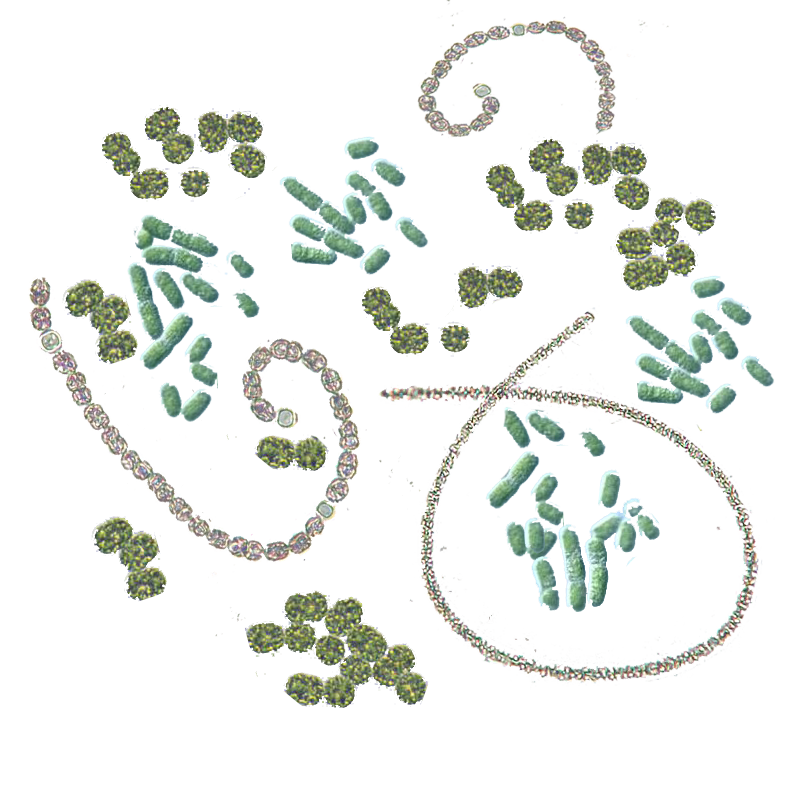
The Cylindrospermopsin Cyanotox qPCR detection kit comes with then new eDNA qPCR hot start mix. This qPCR hot start mix provides excellent performance for sensitive, robust and accurate qPCR detection assay. The eDNA qPCR hot start mix is highly resistant to inhibitory factors, such as humic acids present in environmental samples. This kit contains primers and a probe to detect a highly specific region on the mitochondrial DNA of all species that can produce cylindrospermopsin.
Detection as a service:
We offer the detection of Anatoxin also as a service. See our eDNA service website (in English or in Dutch) or contact us for more information about sampling, sending and pricing.
Kit properties:
The Anatoxin detection kit has:
- High resistance to inhibiting factors. Environmental DNA (eDNA) extracts contain often multiple qPCR inhibiting factors. Normal qPCR master mixes are sensitive to these substances.
- Strong fluorescence signal with low background noise. Isolated environmental samples contain residues of naturally occurring auto fluoresce substances that will interfere with the measurements. Environmental samples needs a strong fluorescence signal from the assay in order not to experience background interference.
- Highest possible sensitivity (1 DNA copy per reaction). Environmental water samples may contain very low amounts of target DNA.
- 100% specificity. Isolated DNA from environmental samples contains billions of DNA fragments from bacteria, protozoa, plants, animals, etc. Not only animals from the same order (fish, amphibians, reptiles, mammals, etc.) must be taken in account during primer/probe design, but all known DNA sequences must be checked for nonspecific binding of the primers and probe.
- ≥ 95% detection probability. The maximum probability of detection can only be achieved with the correct sampling method.
The full method validation report can be found below under “Relevant documents”
The kit has been developed and optimized for use on eDNA isolates purified using the Sylphium Molecular Ecology eDNA Isolation Kit (#SYL002), although other eDNA isolation methods/kits can be used as well. If you need more information on using the obtained isolates from other methods/kits to get reliable results, send an e-mail to info@sylphium.com.
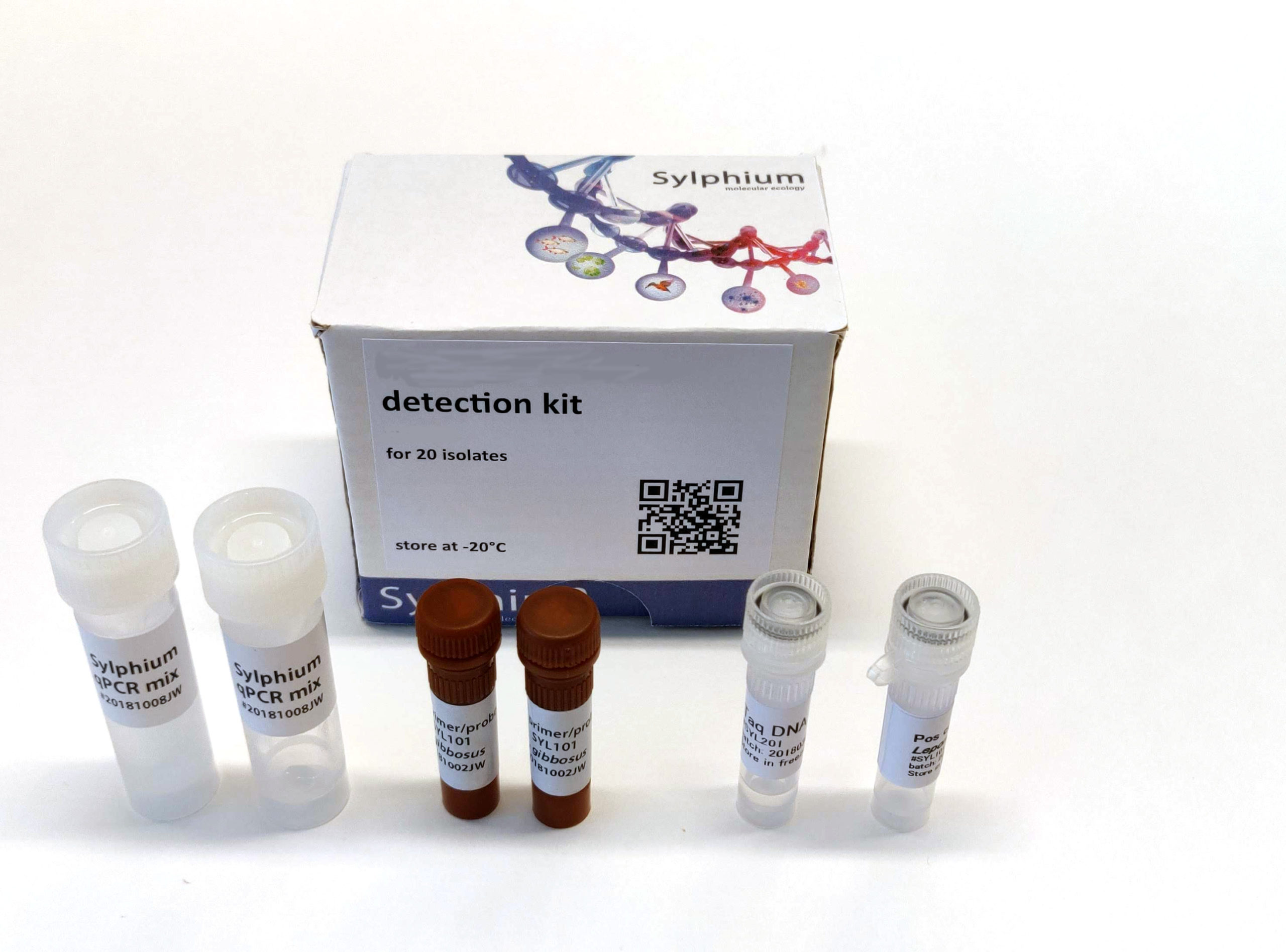
Kit contents:
- Positive control (cloned Cyra gene DNA)
- Primer/probe mix (10x) for detection of cylindrospermopsin (FAM dye)
- eDNA qPCR master mix:
-
- eTaq DNA polymerase
- eTaq qPCR mix
- PCR water
-
The kit contains all materials for 200 reactions of 25µl.
Relevant Documents:
- Technical specifications
- Instructions manual (Universal protocol of eDNA qPCR master mix)
- Method validation report
- Lot validation report: 240418